Preliminary SARS-CoV-2 protease inhibitor shows efficacy in mice
Posted: 20 March 2020 | Victoria Rees (Drug Target Review) | 1 comment
Using X-ray crystallography, researchers have revealed the structure of the SARS-CoV-2 main protease, designing an inhibitor to bind to this target and prevent viral replication.

As part of continued efforts to respond to the COVID-19 pandemic caused by SARS-CoV-2 virus, researchers have used X-ray crystallography to reveal the structure of the SARS-CoV-2 main protease (SARS-CoV-2Mpro) – one of the best characterised drug targets among coronaviruses. They used this information to identify a compound that binds to and inhibits the protease in mice.
With regular news updates and feature articles, our COVID-19 hub has everything you need to keep up to date with R&D during the pandemic. Click below to visit:
…our study provides a useful framework for development of the pyridone-containing inhibitors toward anticoronaviral drugs”
The scientists, from Germany, say their study could drive the improved design of inhibitors to combat the novel coronavirus, a step toward urgently needed therapies to fight the global pandemic. They focused on this protease due to its role in processing polyproteins that are translated from viral RNA.
Based on studying the structure of the main protease, the researchers optimised inhibitors for existing coronaviruses to develop compound 13b, a potent blocker of the SARS-CoV-2 main protease. Using the X-ray structures of SARS-CoV-2 without the ligand, they studied its complex with an α-ketoamide inhibitor derived from a previously designed compound but with the P3-P2 amide bond incorporated into a pyridone ring to enhance the half-life of the molecule in blood.
Drug Target Review has just announced the launch of its NEW and EXCLUSIVE report examining the evolution of AI and informatics in drug discovery and development.
In this 63 page in-depth report, experts and researchers explore the key benefits of AI and informatics processes, reveal where the challenges lie for the implementation of AI and how they see the use of these technologies streamlining workflows in the future.
Also featured are exclusive interviews with leading scientists from AstraZeneca, Auransa, PolarisQB and Chalmers University of Technology.
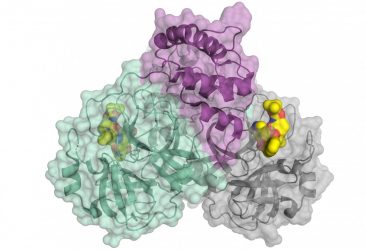

Schematic representation of the coronavirus protease. The enzyme comes as a dimer consisting of two identical molecules. A part of the dimer is shown in colour (green and purple), the other in grey. The small molecule in yellow binds to the active centre of the protease and could be used as blueprint for an inhibitor [credit: HZB].
Lead researcher Linlin Zhang and colleagues found that a high-resolution structure of 13b bound to the protease and blocked viral RNA replication. They report that 13b has features that improve over existing inhibitors, including an extended half-life in blood plasma.
They further tested their leading inhibitor compound in mice, finding that inhalation was well tolerated and that the mice did not show any adverse effects. The researchers suggest that as no human proteases with a similar cleavage specificity are known, this class of inhibitors are unlikely to be toxic.
The team suggest that their results show direct administration of the compound to the lungs may be possible and provide a framework for the development of drugs to combat the novel coronavirus.
“Given these favourable pharmacokinetic results, our study provides a useful framework for development of the pyridone-containing inhibitors toward anticoronaviral drugs,” the authors conclude.
The study was published in Science.
Related topics
Drug Targets, Proteomics, Research & Development, Structural biology, X-ray crystallography
Related conditions
Coronavirus, Covid-19
Related people
Linlin Zhang




Why don’t researchers try a trypsin inhibitor such as cinnamon to inhibit viral replication. It doesn’t have many side effects and apparently trypsin is used by the spike in some way in other related corona viruses to attach to host cells. A number of experiments have shown the effect of trypsin and maybe Cov2 causes the pancreas to increase output.