3D visualisation of COVID-19 surface released for researchers
Posted: 13 March 2020 | Victoria Rees (Drug Target Review) | 21 comments
A new 3D model of the surface of the coronavirus COVID-19 has been released, to aid researchers in the development of a treatment.
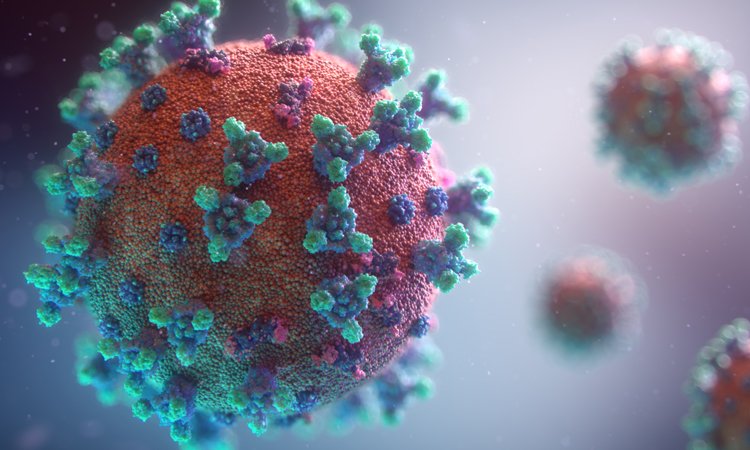

Credit: Fusion Animation
A three-dimensional (3D) model of the surface of the coronavirus COVID-19 has been developed. Created by Fusion Animation, the new model is available for free for scientists to use in the development of treatments to combat the condition.
The model was created by assembling 3D parts together from related COVID-19 coronavirus structures available in public databases. The components used by the developers include:
- Spike (S) protein (PDB code – 6CRV)
- Envelope (E) protein (PDB code – 5X29)
- Membrane (M) protein (PDB code – 3I6G)
They note that the M protein shown is complexed with HLA-A *02 (human leukocyte antigen serotype).
The distribution of these proteins on the surface of the virus was aligned by a random algorithm. The overall representation surface protein density has been reduced to help show S, E and M proteins. The M lipid itself was generated using a particle system to produce a random and organic result.
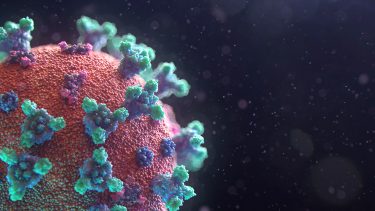

Credit: Fusion Animation
The models were cross-referenced with information from Korkinlab, the US Centers for Disease Control and Prevention (CDC) and the Worcester Polytechnic Institute (WPI), US. The scientists at WPI used a recently published viral genome of the coronavirus made available at the National Center for Biotechnology Information. They then used molecular modelling to reconstruct the 3D structure of major viral proteins and their interactions with human proteins. Their COVID-19 structural genomics map is available to researchers and anyone worldwide.
“We’re confident that our data and visual models could provide the guidance for experimental scientists worldwide who are working feverishly to solve this pandemic,” said Professor Dmitry Korkin, director of the WPI’s bioinformatics and computational biology programme.
According to the developers, the inner structures of the COVID-19 virus are currently being investigated and incorporated into a new visualisation.
Related topics
Disease research, Drug Targets, Imaging, Protein, Proteomics, Research & Development, Structural biology, Targets, Technology, Therapeutics
Related conditions
Coronavirus, Covid-19
Related organisations
Fusion Animation, Korkinlab, US Centers for Disease Control and Prevention (CDC), Worcester Polytechnic Institute



I have read in a file that Corona virus have atleast three viral proteins on its envolope. I would like to know which are them.
Better to target ,to stop the virus entering the host cell primarily,secondly can concentrate to stop the M – protein of the virus,which binds to the host cell.expression of the gene will be same as other cov viruses
Oroxilum indicum, or Indian trumpet flower may inhibit the furin mechanism of virus entry into the cell.
If you consider the envelope to be the lipid membrane of the viron then the three proteins would be M, S and E from most numerous to least.
Is there work going on to determine which specific HLA-A allele expression is more likely to trigger a severe outcome with coronavirus 19 wuhan strain?
Excellent data for researchers and society
Why this viruses don’t effect on birds and animas is that any special structure in there white blood cells
Can the m lipid be dissolved with an aerosol oil such as citrus
It can be dissolved (held in suspension) with soap.
Since this virus is known so far to be man made.and how it does not like things as soap and hand cleaner and breaks down .not to spread .but when it’s in the human body .why try not fighting it with germs that are good.trying something simple as this fight germs against each other.Also known as L. acidophilus or acidophilus, it belongs to the Lactobacillus family of bacteria. Lactic acid bacteria (or L) converts sugars into lactic acid and hydrogen peroxide, substances that inhibit the growth of undesirable bacteria in the intestines
I have researched along the same path and found lipoprotein lipase as the active agent to break down the membrane of Covid-19 and other lipid based viruses. Natural solutions of aerosols delivered into the lungs, which do not harm the alveoli, are the answer. Lipase levels would need to be monitored due to high levels of lipase possibly having a similar effect on the body as seen in pancreatic disfunction. I wish I had a way to test it. Hopefully someone will find something soon.
Thank u very much for illuminating picture of the molecule.
If you have facilities to study the protein structure chemical bonds, you may do work on the same. All such information will help scientists to find the medicine to fight with corona.
Why we cant try inactive virus to create immune response in this case…
And did you analyse cured patient got that immune power or not… It will also helpfull to find medicine for covid 19…
Structure of covoid 19… Please send
It is very informative post and i ask one question which is ” is the coviD 19 attack lecocytes “
What is the molecular structure of the M-lipid? It’s possible that the summer decrease of such viruses is not the heat but the actinic photons. Atmospheric scientists know that free radical oxidants such as OH and HO2, and ozone oxidise lipid surfaces of aerosols as a result of photochemical action. Modelling is needed, but requires the molecular structure of the surface molecules to get the kinetics represented.
I recently realized that after taking the seeds of moringa oleifera my immune system boost and helps fight the illnesses such as cough, fever and covid 19 symptoms and now i feel good. Im not a doctor, please do a research about moringa oleifera chemical contents (coagulant protein, ionizers-polymeres, compounds that founds also in RDS). Something in that plant against covid 19.
Do we know the diameter and/or thickness of the covid-19 lipid sphere?
Do we know the structure of airborne covid-19 as it is released from a shedding host? Are the spheres individually suspended in air, or encased in water droplets suspended in an aerosol, or both?
nice article
It’s there any potential drug or liquid that can weak or dismantling the layer virus of the structure . Virus layer can be distracted from strengthen blood cells or empowered the WBC can weaken the virus. just suggestion.