Interactive 3D model of SARS-CoV-2 surface released
Posted: 25 September 2020 | Victoria Rees (Drug Target Review) | 1 comment
A new interactive map of the surface of SARS-CoV-2, featuring the Spike, Envelope and Membrane proteins, has been released for researchers to use.
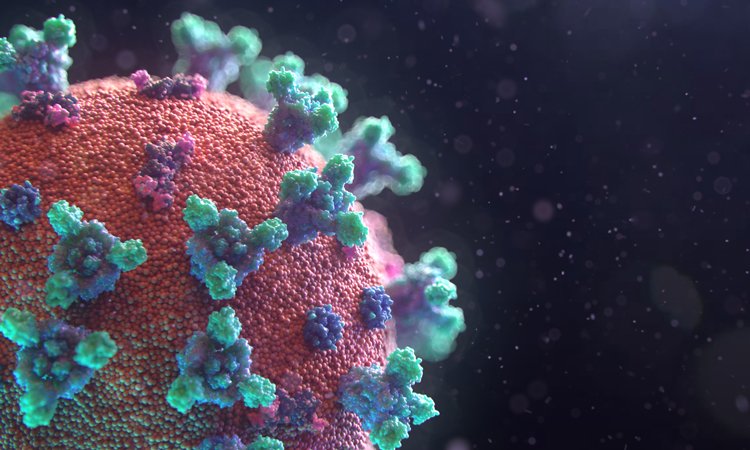

Credit: Fusion Animation.
A new interactive three-dimensional (3D) map of SARS-CoV-2, the virus causing the COVID-19 pandemic, has been released by researchers. The model is free for scientists who wish to use it to aid in the search for a COVID-19 treatment or vaccine.
With regular news updates and feature articles, our COVID-19 hub has everything you need to keep up to date with R&D during the pandemic. Click below to visit:
According to the creators, from Fusion Animation, the model was made by assembling together parts from related 3D coronavirus structures that are available in public databases, such as the RCSB Protein Data Bank (PDB).
Navigation and controls
Move camera: one-finger drag or left mouse button
Pan: two-finger drag or right mouse btton or SHIFT+ left mouse button
Zoom: pinch in/out or mousewheel or CTRL + left mouse button
The developers of the model explain that the components they used are:
- Spike (S) protein (PDB code – 6CRV)
- Envelope (E) protein (PDB code – 5X29)
- Membrane (M) protein (PDB code – 3I6G).
The team highlight that the M protein shown is complexed with HLA-A *02 (human leukocyte antigen serotype).
The creators said that the distribution of these proteins on the surface of the COVID-19 coronavirus was aligned by a random algorithm and the overall representation surface protein density has been reduced to help show the S, E and M proteins. The membrane lipid itself was generated using a particle system to produce a random and organic result.
The researchers also say that the inner structures of SARS-CoV-2 are currently being sourced and incorporated into a new visualisation.
The model was cross-referenced with structures provided by Korkinlab and visual references from the US Centers for Disease Control and Prevention (CDC) and the Worcester Polytechnic Institute (WPI), US.
“We’re confident that our data and visual models could provide the guidance for experimental scientists worldwide who are working feverishly to solve this pandemic,” said Professor Dmitry Korkin, director of the WPI’s bioinformatics and computational biology programme.
Related topics
Disease research, Imaging, Informatics, Protein, Proteomics, Research & Development, Structural biology, Targets
Related conditions
Covid-19
Related organisations
Fusion Animation, Korkinlab, RCSB, US Centers for Disease Control and Prevention (CDC), Worcester Polytechnic Institute




Excellent work! Thanks!