Developing nanobodies and sybodies to combat COVID-19
Posted: 13 November 2020 | Hannah Balfour (Drug Target Review) | No comments yet
Three separate studies have identified nanobodies – a miniature form of antibodies found in camelid species – that can bind to the SARS-CoV-2 Spike (S) protein and neutralise the virus in cells.
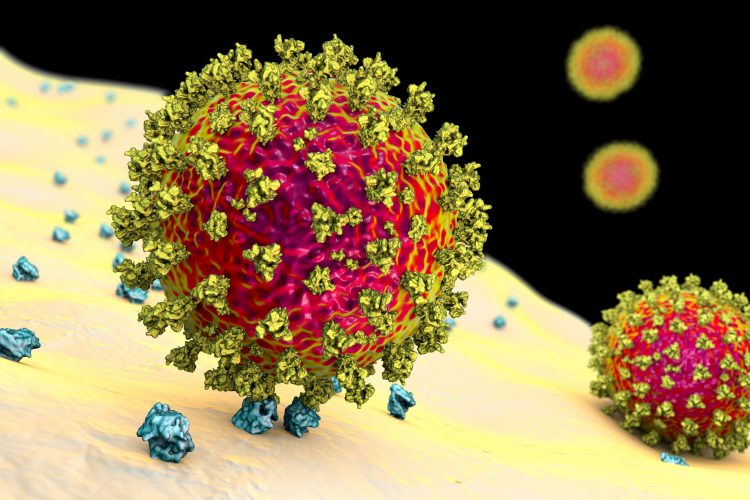

Nanobodies were discovered by chance at the Free University of Brussels in the late 1980s, when students were tasked to extract antibodies from blood and provided with a sample of frozen dromedary serum from the laboratory freezer. When they extracted the antibodies and analysed the results, they found that aside from normal immunoglobulins, the serum contained a previously unseen smaller group of antibodies with no light chains attached. These structures, composed of a single antigen-binding heavy chain fragment, were dubbed nanobodies. Nanobodies have since also been found in other camelid species such as alpacas and llamas.1
With regular news updates and feature articles, our COVID-19 hub has everything you need to keep up to date with R&D during the pandemic. Click below to visit:
![Digram comparing the structures of human antibodies, alpaca heavy chain antibodies and nanobodies [Credit: ChromoTek].](https://drug.russellpublishing.co.uk/wp-content/uploads/Nanobody-versus-antibody-structure-375x131.jpg)
![Digram comparing the structures of human antibodies, alpaca heavy chain antibodies and nanobodies [Credit: ChromoTek].](https://drug.russellpublishing.co.uk/wp-content/uploads/Nanobody-versus-antibody-structure-375x131.jpg)
Digram comparing the structures of human antibodies, alpaca heavy chain antibodies and nanobodies [Credit: ChromoTek].
Due to their smaller size and less complex structure, nanobodies are simpler and cheaper to produce than traditional antibodies. Where full-sized antibodies must be produced in mammalian cells, nanobodies can be produced in bacteria. Another advantage of nanobodies is that they are also more stable. When combined with their small size, this enables multiple potential delivery options, including aerosol-mediated delivery, whereas antibodies are delivered intravenously. These characteristics make them an attractive prophylactic or therapeutic strategy for many diseases, including cancer, multiple sclerosis and novel coronavirus disease 2019 (COVID-19).
Below Drug Target Review’s Hannah Balfour shares three recent studies that describe the discovery and development of nanobodies targeting SARS-CoV-2, the virus that causes COVID-19.
Drug Target Review has just announced the launch of its NEW and EXCLUSIVE report examining the evolution of AI and informatics in drug discovery and development.
In this 63 page in-depth report, experts and researchers explore the key benefits of AI and informatics processes, reveal where the challenges lie for the implementation of AI and how they see the use of these technologies streamlining workflows in the future.
Also featured are exclusive interviews with leading scientists from AstraZeneca, Auransa, PolarisQB and Chalmers University of Technology.
The discovery of sybody 23
In the first study,2 researchers designed and selected a synthetic nanobody, called a sybody, to target the receptor binding domains (RBDs) of the SARS-CoV-2 Spike (S) protein. SARS-CoV-2 depends on interactions between the S protein on its surface and the human cell surface protein angiotensin converting enzyme 2 (ACE2) to infect cells. The S protein binds ACE2 using three RBDs, which look like finger-like protrusions. Because SARS-CoV-2 is dependent on this interaction to infect cells, researchers believe that preventing it could prevent the spread of the virus.
To identify the best sybody candidates from existing large synthetic libraries, Christian Löw’s group from the European Molecular Biology Laboratory (EMBL) in Hamburg, Germany, used a technology platform that was recently developed in the lab of Markus Seeger at the University of Zurich, Switzerland.
Using the S protein RBDs as bait, the team were able to identify several sybodies that could target SARS-CoV-2. They then tested these sybodies to establish their stability, how precisely they bound to the RBDs and their effectiveness at blocking the domains. One candidate, called sybody 23, was found to be particularly effective in blocking the RBDs.
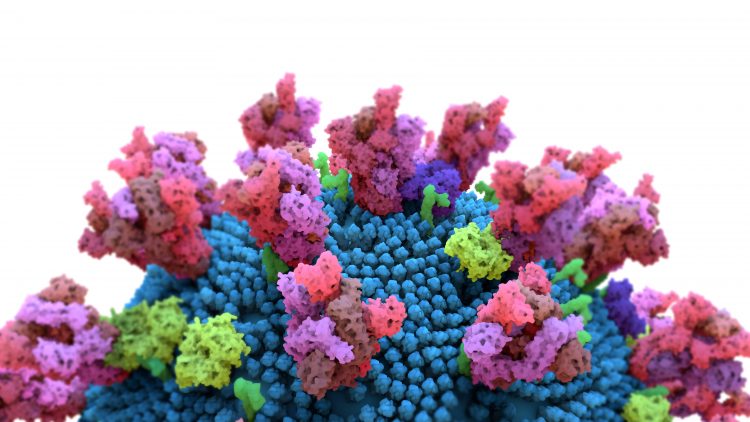

To understand exactly how sybody 23 interacts with the viral RBDs, researchers in the group of Dmitri Svergun at EMBL Hamburg analysed the binding of sybody 23 to the RBDs using small-angle X-ray scattering. Martin Hällberg at the Center for Structural Systems Biology (CSSB) in Germany and Karolinska Institutet in Sweden also used cryogenic electron-microscopy (cryo-EM) to determine the structure of the full SARS-CoV-2 S protein bound to sybody 23.
The SARS-CoV-2 RBDs switch between two positions – up and down. Up is the position required to interact with ACE2 and initiate viral entry, as the RBDs are accessible. In the down position the RBDs are inaccessible, furled in to protect against detection by the immune system. According to the team, sybody 23 can bind to either, blocking the areas where ACE2 would normally bind. They suggest that its ability to block RBDs regardless of their position might explain why sybody 23 is so effective.
In a final experiment, Ben Murrell and his researchers at Karolinska Institutet used a lentivirus modified to express the SARS-CoV-2 S protein to establish whether sybody 23 can neutralise SARS-CoV-2. They observed that sybody 23 successfully disabled the modified virus in vitro but cautioned that additional testing would be required to confirm whether sybody 23 could stop SARS-CoV-2 infection in the human body.
Developing a trivalent nanobody to neutralise SARS-CoV-2
In the second study,3 researchers screened a library of synthetic nanobody sequences displayed on the surface of yeast to identify 21 candidates that could prevent the SARS-CoV-2 S protein from interacting with ACE2. Using cryo-EM, researchers from the University of California at San Francisco, Icahn School of Medicine at Mount Sinai and other locations, all US, were able to show that one nanobody, Nb6, binds the S protein in a fully inactive conformation, where its RBDs are locked into their inaccessible down-state, rendering the virus incapable of binding ACE2.
From the pool of nanobodies able to bind the S protein, the team developed a trivalent nanobody, which they named mNb6-tri. According to their study, mNb6-tri has femtomolar affinity for the S protein and is extremely potent, able to neutralise SARS-CoV-2 at picomolar concentrations. The trivalent nanobody was also shown to retain its function after aerosolisation, lyophilisation and heat treatment, so it could be delivered directly to the epithelial cells impacted by SARS-CoV-2 infection using aerosol-mediated delivery.
“The combined stability, potency and diverse epitope engagement of our nanobodies… provide a unique potential prophylactic and therapeutic strategy to limit the continued toll of the COVID-19 pandemic,” wrote the authors of the paper.
Wally the llama and his nanobodies
In the final paper4 researchers at the University of Pittsburgh (Pitt) School of Medicine in the US, describe how to extract SARS-CoV-2 antibody fragments from llamas.
![Photo of a black llama wearing a red headcollar - Wally the Llama [Credit: Sonya Paske, Capralogics Ltd.].](https://drug.russellpublishing.co.uk/wp-content/uploads/nanobody-llama-167x250.jpg)
![Photo of a black llama wearing a red headcollar - Wally the Llama [Credit: Sonya Paske, Capralogics Ltd.].](https://drug.russellpublishing.co.uk/wp-content/uploads/nanobody-llama-167x250.jpg)
Wally the Llama [Credit: Sonya Paske, Capralogics Ltd.].
In the study, the researchers describe how they immunised a llama called Wally with a piece of the SARS-CoV-2 S protein. After two months they collected blood from Wally and used a cryo-EM technique that the paper’s senior author, Dr Yi Shi, Assistant Professor of Cell Biology, had been perfecting to identify the mature SARS-CoV-2 S protein neutralising nanobodies.
Then, with the help of Pitt’s Center for Vaccine Research (CVR), the scientists exposed their nanobodies to live SARS-CoV-2 virus and found that only a fraction of a nanogram was sufficient to neutralise enough virus to spare a million cells from being infected. According to the team, this makes their nanobodies some of the most effective therapeutic antibody candidates for SARS-CoV-2, hundreds to thousands of times more effective than other nanobodies that have been discovered before.
Aside from their incredible potency, the researchers also explained that the stability of their nanobodies make them a highly desirable therapeutic option. In their paper they demonstrate that these nanobodies could sit for at least six weeks at room temperature and tolerate being fashioned into an inhalable mist without loss of efficacy.
Shi explained that because traditional SARS-CoV-2 antibodies require intravenous administration, diluting the product throughout the body, a much larger dose is needed which comes with a higher price tag. “Nanobodies could potentially cost much less,” he said. “They are ideal for addressing the urgency and magnitude of the current crisis.”
By collaborating with Pitt’s Dr Cheng Zhang and Dr Dina Schneidman-Duhovny from the Hebrew University of Jerusalem in Israel, the researchers were able to show that their nanobodies use a variety of mechanisms to block SARS-CoV-2 infection. The team suggest this means that, as in the study above, they may be able to bioengineer various nanobodies and create ones that are linked together with multiple mechanisms of action, thereby preventing SARS-CoV-2 from developing resistance.
“As a virologist, it is incredible to see how harnessing the quirkiness of llama antibody generation can be translated into the creation of a potent nanoweapon against clinical isolates of SARS-CoV-2,” said study co-author and CVR Director, Dr Paul Duprex.
References
- Discovery of Nanobodies [Internet]. ChromoTek GmbH. Available at: https://www.chromotek.com/technology/discovery-of-nanobodies/
- Custódio, T.F., Das, H., Sheward, D.J. et al.Selection, biophysical and structural analysis of synthetic nanobodies that effectively neutralize SARS-CoV-2. Nat Commun 11, 5588 (2020). https://doi.org/10.1038/s41467-020-19204-y
- Schoof, M., Faust, B., Saunders, R., et al. An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive Spike. Science. 2020;:eabe3255. DOI: 10.1126/science.abe3255
- Xiang, Y., Nambulli, S., Xiao, Z., et al. Versatile and multivalent nanobodies efficiently neutralize SARS-CoV-2. Science. 2020;:eabe4747. DOI: 10.1126/science.abe4747
Related topics
Antibodies, Biologics, Drug Development, Drug Targets, Imaging, In Vitro, Protein, Proteomics, Research & Development, Screening, Therapeutics
Related conditions
Coronavirus, Covid-19
Related organisations
Center for Structural Systems Biology (CSSB), European Molecular Biology Laboratory (EMBL) in Hamburg, Icahn School of Medicine at Mount Sina, Karolinska Institutet, The Hebrew University of Jerusalem, University of California at San Francisco, University of Pittsburgh School of Medicine, University of Zurich (UZH)



