Researchers map vital atomic-scale protein on COVID-19
Posted: 20 February 2020 | Victoria Rees (Drug Target Review) | 10 comments
Using cryogenic electron microscopy, a team has mapped the Spike protein on COVID-19, which could be used in the development of vaccines.


Scientists have announced a critical breakthrough in their research for a vaccine to comabt COVID-19 (coronavirus) by creating the first three-dimensional (3D) atomic-scale map of the section of the virus that attaches to and infects human cells.
With regular news updates and feature articles, our COVID-19 hub has everything you need to keep up to date with R&D during the pandemic. Click below to visit:
Conducted by the University of Texas (UT) at Austin and the National Institutes of Health (NIH), both US, the study has produced a map of the Spike (S) protein, which may allow researchers to design and make vaccines and antiviral drugs. The researchers are also now working on a related viable vaccine candidate stemming from their research.
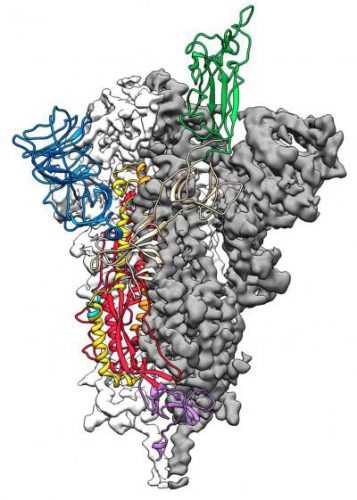

This is a 3D atomic scale map, or molecular structure, of the 2019-nCoV Spike protein. The protein takes on two different shapes, called conformations – one before it infects a host cell and another during infection. This structure represents the protein before it infects a cell, called the prefusion conformation (credit: Jason McLellan/UT at Austin).
The team used cryogenic electron microscopy (cryo-EM) to make their atomic-scale 3D model. The molecule the team produced – and for which they obtained a structure – represents only the extracellular portion of the S protein, but is enough to elicit an immune response in patients.
Drug Target Review has just announced the launch of its NEW and EXCLUSIVE report examining the evolution of AI and informatics in drug discovery and development.
In this 63 page in-depth report, experts and researchers explore the key benefits of AI and informatics processes, reveal where the challenges lie for the implementation of AI and how they see the use of these technologies streamlining workflows in the future.
Also featured are exclusive interviews with leading scientists from AstraZeneca, Auransa, PolarisQB and Chalmers University of Technology.
Jason McLellan, associate professor at UT Austin who led the research, and his colleagues have spent many years studying other coronaviruses, including SARS-CoV and MERS-CoV. They had already developed methods for locking coronavirus S proteins into a shape that made them easier to analyse and could effectively turn them into candidates for vaccines. This experience gave them an advantage for studying the novel virus.
“As soon as we knew this was a coronavirus, we felt we had to jump at it,” McLellan said, “because we could be one of the first ones to get this structure. We knew exactly what mutations to put into this, because we’ve already shown these mutations work for other coronaviruses.”
Two weeks after receiving a genome sequence of the virus from Chinese researchers, the team designed and produced samples of their stabilised S protein. It took approximately 12 more days to reconstruct the 3D atomic scale map of the S protein and submit a manuscript.
Next, McLellan’s team plans to use their molecule to attack the virus that causes COVID-19, using it as a ‘probe’ to isolate naturally produced antibodies from patients who have been infected with the virus and successfully recovered. In large enough quantities, these antibodies could help treat a coronavirus infection soon after exposure.
The study was published in Science.
Related topics
Drug Targets, Imaging, Microscopy, Protein, Structural biology, Targets, Vaccine
Related conditions
Covid-19
Related organisations
University of Texas (UT) at Austin, US National Institutes of Health (NIH)
Related people
Jason McLellan




Are you disclosing the fact that the 3 sequence insertions in the otherwise exact copy of SARS spike protein are precisely at the binding site with the ACE2 receptor?
This is a brilliant study, hopefully helpful also for solving the practical issues of the highly contagious Coronavirus spread.
It certainly is elucidating to view the 3D structure of COVID; but where can one find the definitive nucleotide sequence of the SPIKE PROTEIN of COVID ? That is requisite for the recombinant production of the SPIKE PROTEIN !
Dr . Mike Huchital, DYNE IMMUNE INSTITUTE
here is the entire genome: https://www.ncbi.nlm.nih.gov/nuccore/NC_045512
the S protein is YP_009724390.1 . Im not sure about the the other strain, as they are supposedly 2
S and L protein
Hi folks, Is there a screening system for Covid-19 to check the activity of small molecule/peptide? OR We have screened using luciferase activity? targeting PLPro or 3CLpro. We could not find a screening system. If you share your views regarding the screening system it will be highly appreciated. Thank you,
For breaking glycogen protein and devlop antibodies for protect wbc and immunity brek structure of covid 19
Attachment is in thing, affinity is another. What is the net charge of the structure. Aytipical lymphs, large granular lymphs noted in patients. I’ve been exposed at least 3 or 4 times to this virus starting in November, treatment Choroseptic throat sprAy to prevent discrimination from upper to lower respiratory tract. Phenols
S protein cleavage occurs at two sites within the S2 portion of the protein. Fusion generally occurs within ACIDIFIED endosomes. Can we use a regular sodium carbonate Na2CO3 to for prevention purpose and possibly for some treatment as acid neutralizer agent ?
Hi, is there a Dalton mass number for this protein or parts of it? Much appreciated.