Biomarkers as molecular footprints for cancer therapy: current perspectives and promises
Posted: 3 December 2015 | Firoz Anwar (King Abdulaziz University), Mahfoozur Rahman (Sam Higginbottom Institute of Agriculture Technology & Sciences), Sarwar Beg (Panjab University), Vikas Kumar (Sam Higginbottom Institute of Agriculture Technology & Sciences) | 1 comment
Knowledge about cancer biomarkers has increased tremendously and provides clinicians and researchers with valued opportunities to understand the molecular mechanism of cancer, thus applying it to effective, early-stage diagnosis and treatment. Cancer biomarkers include a vast range of biochemical molecules such as nucleic acids, proteins, sugars, lipids, small metabolites, cytogenetic and cytokinetic parameters, as well as whole tumour cells inside the body. A comprehensive understanding of each biomarker enables the perfect diagnosis of the disease and makes it easy to choose from multiple different therapeutics, although it is important to note that technology cannot currently identify all biomarkers that play a key role in cancer. This article provides an account of several biomarkers used in the diagnosis, prognosis and treatment of cancer, discussing the uses of some of the markers that are both already in clinical practice, and in trials.

Increased knowledge of cancer biology, combined with rapid advancements in molecular technology, have led to the recognition of umpteen numbers of biomarkers in cancer. To that end, a paper is published almost on a daily basis in this realm1,2. The availability of abundant literature makes the task easier for clinicians and scientists to critically understand the biomarkers, which helps in understanding how they can be targeted and incorporated into routine medical practice3,4. The National Cancer Institute defines biomarkers as footprint molecules found in blood, other body fluids and tissues, which flag up normal or abnormal process of any disease condition5. Every cell has its own unique molecular signature, which gives rise to certain activities by the response of genes and proteins5,6. Therefore, evaluation of biomarkers is an important approach for understanding the biological and pathogenic processes or pharmacological responses for therapeutic intervention7.
Every year more than 11 million people are diagnosed with cancer and it is estimated that the number will reach up to 16 million by 20207. A complex disease, cancer involves multistep processes involving conversion of proto-oncogenes to oncogenes, which may lead to the development of unmatured group of cells (tumours)8. There are three main categories of genes involved in the development of cancer: proto-oncogenes, tumour suppressor genes and DNA repair genes8. Epigenetic changes such as DNA methylation and changes in pattern of histone modification can also lead to cancer, and such changes may result in further alteration in chromatins, leading to abnormal gene expression9. Modern technology can be used to understand the footprints of cells which become cancerous, and their recognition is important for early cancer diagnosis, improved patient prognosis and therapy selection9. Using biomarkers, it is easy to locate the tumour and determine its stage and subtypes for undertaking necessary therapeutic interventions.
Genetic cell alteration, which includes gene rearrangements, point mutations and gene amplifications, is profound in cancer cells. These may alter several molecular pathways, regulating cell growth and metastasis10-11. To date, genetics, genomics, proteomics and many non-invasive imaging techniques allow several biomarkers to be measured11, and scientists now have a broad understanding of disease pathways, corresponding protein targets and pharmacotherapy for various disorders beyond cancer. Nowadays, the major challenges in cancer are the difficulties in establishing the exact relationship between cancer biomarkers and the clinical pathology, and non-invasively detecting the tumour at an early stage11. Similarly, recognition of small changes in genomics and proteomics can highlight molecular targets for developing cancer therapeutics.
Drug Target Review has just announced the launch of its NEW and EXCLUSIVE report examining the evolution of AI and informatics in drug discovery and development.
In this 63 page in-depth report, experts and researchers explore the key benefits of AI and informatics processes, reveal where the challenges lie for the implementation of AI and how they see the use of these technologies streamlining workflows in the future.
Also featured are exclusive interviews with leading scientists from AstraZeneca, Auransa, PolarisQB and Chalmers University of Technology.
Diagnostic and prognostics biomarkers are useful for clinical oncologists, since they aid in identifying the suspected patients who are at a high risk for the disease and in diagnosing them at an early stage12. These biomarkers are available in various forms, such as traditional biomarkers, which can be easily assessed with radiological techniques such as mammography, and detection of tumour-specific antigens (e.g., prostate-specific antigen; PSA)12. The availability of the complete human genome and advancement in technologies such as high-throughput DNA sequencing, microarray and mass spectrometry, have, in this regard, made the task tremendously easier8,12. The list of cancer biomarkers have dramatically expanded to include the sequence and expression level of DNA, RNA and protein, as well as metabolites12. Cancer biomarkers are depicted in Figure 1.
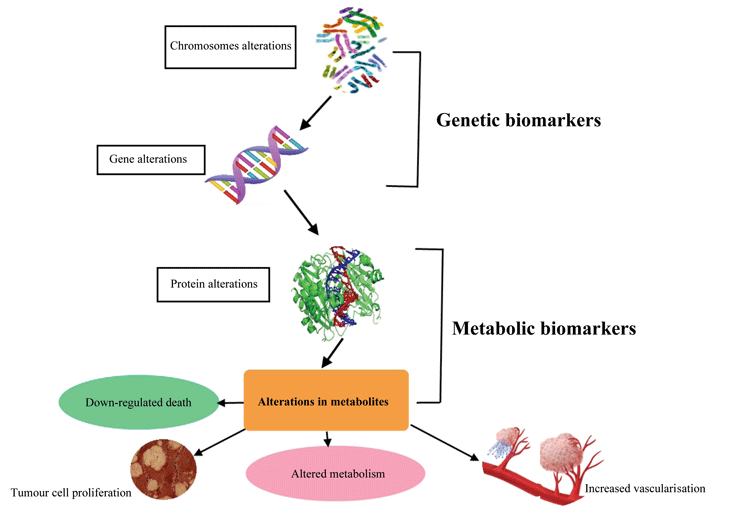

Figure 1: Pictorial depiction of various cancer biomarkers
Examples of of biomarkers in cancer
Cells as biomarkers
In the advanced stage of cancer, cells start appearing in the blood stream. Circulating tumour cells (CTCs) are recognised as powerful biomarkers in oncology and have been predicted in metastatic breast cancer. They provide an early, reliable indication of disease progression and survival for patients on systemic therapy for metastatic breast cancer. Therefore, elimination of CTCs could be a better approach in breast cancer therapy13. This is summarised in Table 1 for different diseases.
Table 1: Biomarkers in cancer
|
Cancer Biomarker |
Types of tumour |
Reference |
|
Metabolic biomarkers |
||
|
Glucose Metabolism |
All types of cancer covered |
[4, 5] |
|
Cells as biomarker |
||
|
Circulating tumour cells (CTCs) |
Metastatic breast cancer |
[8-10] |
|
Cancer stem cell |
Brain tumour, breast cancer and prostate cancer |
[8-10] |
|
Biomolecules as biomarker |
||
|
Prostate specific antigen |
Prostate cancer |
[12, 13] |
|
Alpha-fetoprotein |
Hepatocellular carcinoma |
[7, 10] |
|
BRCA-1, BRCA-2 |
Colorectal cancer |
[11, 12] |
|
Heat shock proteins (Hsp27 and Hsp70) |
Cervical, uterine, and prostate carcinoma |
[13, 14] |
|
Throglobulin |
Papillary and thyroid cancer |
[8, 15] |
|
Cancer antigen 125 |
Ovarian cancer |
[11-13] |
|
Human chorionic- gonadotrophin (hCG) |
Ovarian and testicular cancer |
[6, 11] |
|
Genetic biomarkers |
||
|
Phiadelphia chromosomes and Bcl-2 gene |
Burkitt lymphoma |
[9, 11] |
|
APC gene |
Adeno-carcinoma and squamous cell carcinoma |
[10-13] |
Cytogenetic and cyto-kinetic markers
Any changes in chromosomes are classical markers of cancer, since the association between chromosomal aberration and neoplastic transformation has been well established. For example, aneuploidy has frequently been found in malignant tumours. Somatic mutations, which are associated with malignant transformation, are promising biomarkers for cancer risk. Meanwhile, genome-based biomarkers have been identified in certain epithelial tumours such as mRNA of cytokeratin-19, EGFR and MU-1, etc10,13. Recently, exon-3 has been observed as a novel transcriptome marker, which is primarily involved in prostate cancer cell proliferation and endocrine cancer. Elsewhere, protein encoded by the mini-chromosome maintenance gene has emerged as useful marker for cell proliferation10,13.
Viral biomarkers
Hepatocellular carcinoma (HCC) is one of the most common cancers worldwide and a major cause of death in developing countries. Infection with either hepatitis B virus or hepatitis C virus, both of which promote carcinogenesis of HCC, is known to be a major cause of HCC14. Another cancer in which viral biomarkers can be implicated is cervical cancer. As the second most widespread gynecological cancer in women, persistent infection by human papillomaviruses (HR-HPVs) is known to be the major cause of the main types of cervical cancer – squamour cell cancer and adenocarcinoma, with other risk factors such as smoking and use of the pill also acting as risk factors. HPVs are also profound in oral, oesophageal and vaginal, vulvar and penile cancer14-15.
Genetic biomarkers
Cancer is also mediated by alterations in oncogene and tumour suppressor genes, which regulates cell proliferation and other homeostatic functions7. Non-random mutations, translocation, and rearrangement in the regulatory region of chromosomes and genes are also known to be associated with particular types of malignancy. For example, the ‘Philadelphia chromosome’ associated with chronic myelogenous leukaemia results from translocation between chromosomes 9 and 22, leading to cancer-causing gene BCR-ABL7,16,17. Meanwhile, several tumour suppressor genes, such as p53, Rb, DCC, Brush-1, BRCA-1, and BRCA-2, have been implicated in breast tumours7,16,17.
Conclusion
Cancer is a complex disease; molecular heterogeneity and adaptive resistance makes it challengeable, thus its exact pathomechanism needs be understood for patients to receive effective chemotherapy. Cancer biomarkers have paved the way for the development of novel pharmaceuticals and, therefore, it is vital that clinical, translational and laboratory-based researchers become aware of the issues related to biomarker development.
References
- Srinivas PR, Kramer BS, Srivastava S. Trends in biomarker 1. Research for cancer detection. Lancet Oncol 2001; 2: 698- 704
- Cho WSC. Contribution of onco-proteomics to cancer 2. Biomarker discovery. Mol Cancer 2007; 6: 25
- Hanahan D, Weinberg RA. The hallmarks of cancer.3. Cell 2000; 100: 57-70
- Bayli SB, Ohm JE. Epigenetic gene silencing in cancer-a. Mechanism for early oncogenic pathway addiction? Nature Rev Cancer 2006; 6: 107-17
- Ludwig JA, John N. Weinstein biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer 2005; 5: 845-56
- Sidransky BD. Emerging molecular markers of cancer. Nat Rev Cancer 2002; 2: 210-9
- Vogelstein CB, Kinzler KW. Cancer genes and the pathways they control. Nat Med 2004; 10: 789-99
- Rahman M, Ahmad MZ, Kazmi I, et al. Advancement in multifunctional nanoparticles for the effective treatment of cancer. Expert Opin Drug Deliv. 2012; 9 (4): 367-81
- Sawyers CL. The cancer biomarker problem. Nature 2008; 452: 548-52
- Whitfield ML, George LK, Grant GD, Perou CM. Common markers of proliferation. Nat Rev Cancer 2006; 6: 99-106
- Toru H, Masaharu Y, Shinji T, Kazuaki C. Genetic polymorphisms and head and neck cancer risk. Int J Oncol 2008; 32: 945-73
- Shinjo K, Kondo Y. TargetingCancer Epigenetics: Linking Basic Biology to Clinical Medicine. Adv Drug Deliv Rev. 2015
- Frantzi M, Latosinska A, Merseburger AS, Mischak H. Recent progress in urinary proteome analysis for prostatecancer diagnosis and management. Expert Rev Mol Diagn. 2015; 22:1-16
- Middleton FK, Patterson MJ, Elstob CJ. Commoncancer-associated imbalances in the DNA damage response confer sensitivity to single agent ATR inhibition. Oncotarget. 2015; 6(32): 32396-409
- Rahman M, Akhter S, Ahmad MZ, Ahmad J, Addo RT, Ahmad FJ, Pichon C. Emerging advances in cancer nanotheranostics with graphene nanocomposites: opportunities and challenges. Nanomedicine (Lond). 2015; 10 (15): 2405-22
- Rahman M, Ahmad MZ, Kazmi I, Akhter S, Afzal M, Gupta G, Sinha VR. Emergence of nanomedicine as cancer targeted magic bullets: recent development and need to address the toxicity. Curr Drug Discov Technol. 2012; 9(4): 319-29
- Ahmad MZ, Akhter S, Jain GK, Rahman M, Pathan SA, Ahmad FJ, Khar RK. Metallic nanoparticles: technology overview & drug delivery applications in oncology. Expert Opin Drug Deliv. 2010; 7(8): 927-42
Biographies
Mr. Mahfoozur Rahman* is a young Research Scientist. Presently, he works as a research fellow at the Department of Pharmaceutical Sciences, Faculty of Health Sciences, Sam Higginbottom Institute of Agriculture, Technology & Sciences, in Allahabad, India, under the supervision of Dr. Vikas Kumar. He is intensively involved in the development of nano-sized drug delivery systems for the treatment of inflammatory disorders. He has been selected in UGC networking for his research work in the liposome research laboratory, UIPS, at Panjab University in Chandigarh, India. He is a member of IPGA and editorial board member of various journals. He has an international book chapter in CRC press (Taylor and Francis), Springer and Elsevier and published his research findings in various peer-reviewed journals of international repute. *E-mail: mahfoozkaifi@gmail.com
Mr Sarwar Beg is a UGC-Doctoral Meritorious Research Fellow in Science at University Institute of Pharmaceutical Sciences, Panjab University, in Chandigarh, India. He completed his Masters in Pharmaceutics from Hamdard University in New Delhi. His major areas of research interest include DoE/QbD-based development and characterisation of controlled-release drug delivery systems, and bioenhanced drug delivery systems like self-nanoemulsifying systems, lipidic nanoparticles, liposomes, microspheres, nanoparticles, dendrimers, carbon nanotubes and nanocomposites. To date he has authored more than 30 publications in various high impact peer-reviewed journals, 10 book chapters, two books and has one Indian patent to his credit.
Dr. Vikas Kumar*, Assistant Professor (Pharmacognosy), Department of Pharmaceutical Sciences, Sam Higginbottom Institute of Agriculture, Technology & Sciences, Allahabad, India, has been working on traditional medicine in drug discovery leading to the development of therapeutic leads from natural resources. His research work is highlighted on screening, evaluation, formulation and standardisation of herbal drugs with their validation to ensure quality, efficacy and safety. He has made innovative, outstanding and original contributions both in research and education in the area of natural products. He has to his credit above 30 publications in peer-reviewed impact journals, three books and three book chapters. *E-mail: phvikas@gmail.com
Prof Firoz Anwar is working as Professor, Department of Biochemistry, Faculty of Science, King Abdulaziz University in Jeddah, Saudi Arabia. He has been working on screening, evaluation, formulation and standardisation of herbal drugs as well as synthetic drugs. He was the chairman and Ex Dean of Siddhartha Institute of Pharmacy, Dehradun, India. Dr. Firoz Anwar is a pharmacist and completed his master and PhD from Jadavpur University. His research career has been outstanding, including globally acclaimed contributions to development from natural resources including Ayurveda, ethnopharmacology, herbal drug technology and synthetic drugs. His pioneering work has led to many important national and international projects in the field of pharmacology and clinical pharmacology. Based on these works, he has to his credit above 100 publications in peer-reviewed journals, several patents, books and chapters.
Related topics
Biomarkers, Gene Therapy, Oncology
Related organisations
King Abdulaziz University, Panjab University, Sam Higginbottom Institute of Agriculture Technology & Sciences
Related people
Firoz Anwar, Mahfoozur Rahman, Sarwar Beg, Vikas Kumar




I would like to invite the authors to review the work of Dr. Bernard A Fox, CEO of UbiVac, and recently recognized by the Society for Immunotherapy of Cancer for his contributions to this international organization. I will be pleased to make an introduction.