Human iPSC-derived functional cells are revolutionising phenotypic drug discovery and development
Posted: 4 September 2018 | Arie Reijerkerk, Farbod Famili, Mieke Doornbos, Stefan Braam | No comments yet
Effective drug discovery and development relies largely on the availability of predictive pre-clinical model systems; the absence of which has contributed to late-stage drug failures and high expenditures. New industry acceptance of human induced pluripotent stem cell (hiPSC)-derived cardiomyocyte assay systems for regulatory safety pharmacology is now resulting in broad use of the technology in drug discovery and development. Here, we demonstrate an end-to-end solution covering large-scale manufacturing of hiPSC-derived cells enabling phenotypic high-throughput screening (HTS) in a relevant human cellular disease model.

Despite many technological developments over the past decades, the output of commercial drug discovery and development has been relatively ineffective; which has been referred to as the productivity crisis in pharmaceutical industry by many experts.1,2 The reality is that this crisis makes the modern drug development process very expensive and the costs have shown a significant upward trend for decades. It was estimated that the accumulated costs per approved new compound are in the region of $2,558 million. Interestingly, $1,098 million of this is spent in the pre-clinical development phase. A significant contributor to these high costs per approved drug is the extremely low success rates in clinical drug development; only 11.8% of drug candidates entering Phase I are likely to be approved.3 This number is even lower (~1%) if the whole drug development process, from target discovery to market launch, is taken into account.
The reasons for the current low levels of R&D productivity are complex, but some factors have been identified. Historically, drugs have been successfully developed using a low-throughput approach comprising ‘phenotypic’ testing of a small set of molecules in intact animals to assess the therapeutic and pharmacological properties of each molecule. However, in recent decades the elucidation of the human genome and technological developments in, for example, chemistry have led to the implementation of a more target-based drug discovery approach centring around a selected protein or signalling pathway in cells. Although simultaneous developments in high-throughput technologies enabled the screening of large compound libraries, the potential for success of target-driven approaches has been abolished by the incorrect assumption of a direct link between modulation of the selected target and the potential to treat or mitigate the disease. In addition, target-based drug screening generally uses a combination of artificial cell and animal models, in fact not representing the complexity of human (disease) biology (Figure 1, top). Building true confidence that modulation of these targets is indeed relevant for the human disease typically requires development of the drug up to the costly phase 2/3 clinical trials stage. Nowadays, the drawbacks of target-based drug discovery are being increasingly recognised by pharma industries, which has led to a renewed interest in the use of phenotypic screening approaches.2,4,5
This was catalysed by the thought-provoking outcome of a meta-analysis of drug discovery approaches from 2011 showing that most of the latest newly-developed first-in-class medicines (ie, completely novel drugs) were actually obtained using a phenotypic drug discovery approach and the minority originated from target-based discovery programmes.6 Between 2000 and 2008, of the 50 first-in-class small molecule drugs discovered, 28 originated from phenotypic strategies compared to 17 from the target-based approach. This has a led to a strong desire to build human disease-relevant phenotypic assays (ie, those with a clear link to human disease in combination with functional readouts closely related to the desired clinical readout in patients) that can be utilised for both high-throughput screening and further development of the molecule of interest.
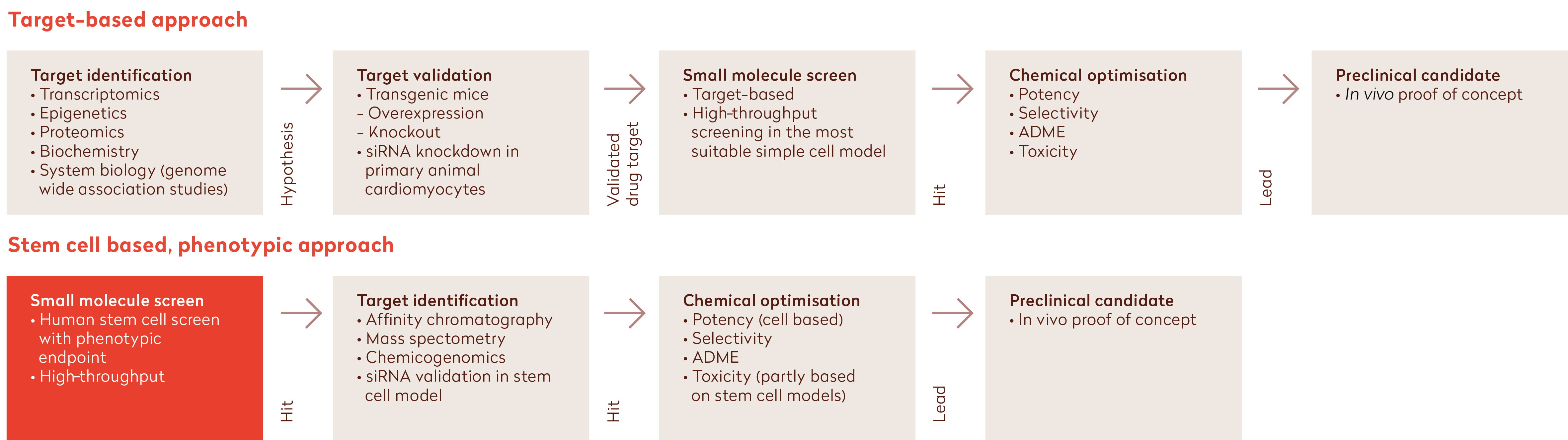

Figure 1: Comparison of the target-based approach (top) with the stem cell-based phenotypic approach (bottom) in drug discovery (adapted from12). Introducing hiPSC-derived cellular models into the drug discovery screening process through high-throughput screening using relevant human assay systems enables reliable efficacy testing of drugs and results in lower attrition rates compared to target-based approaches.
iPSC technologies are revolutionising phenotypic drug discovery and development
The benefit of phenotypic screening has been attributed to its unbiased nature, which allows for ‘serendipity’ and is not necessarily hampered when the disease process is not (yet) fully understood. Although the availability of certain types of primary human cells and (patient-derived) tissues required for phenotypic screening has increased remarkably in recent years, application of human cellular models from tissues that are difficult to access, such as cardiomyocytes and neurons, remains challenging. The availability of human induced pluripotent stem cell (hiPSC) technologies holds great promise to overcome this challenge and improve decision-making in the preclinical phase, which ultimately should reduce the time and cost of bringing new drugs to the market (Figure 1, bottom). In fact, diseased iPSC platforms have already yielded the first clinical candidates.7 Using a patient-specific iPSC approach, GlaxoSmithKline successfully repurposed their Retigabine, which is currently tested in a phase II clinical trial for Amyotrophic Lateral Sclerosis. Scientists at companies such as Ncardia are developing end-to-end hiPSC-based (disease) assay screening solutions for drug discovery and development, which represent the complexity of human biology (Figure 2). With this they provide scalable and predictive solutions to measure compound efficacy and safety at high throughput, thereby improving the decision-making process and ultimately reducing the overall drug discovery and development costs.
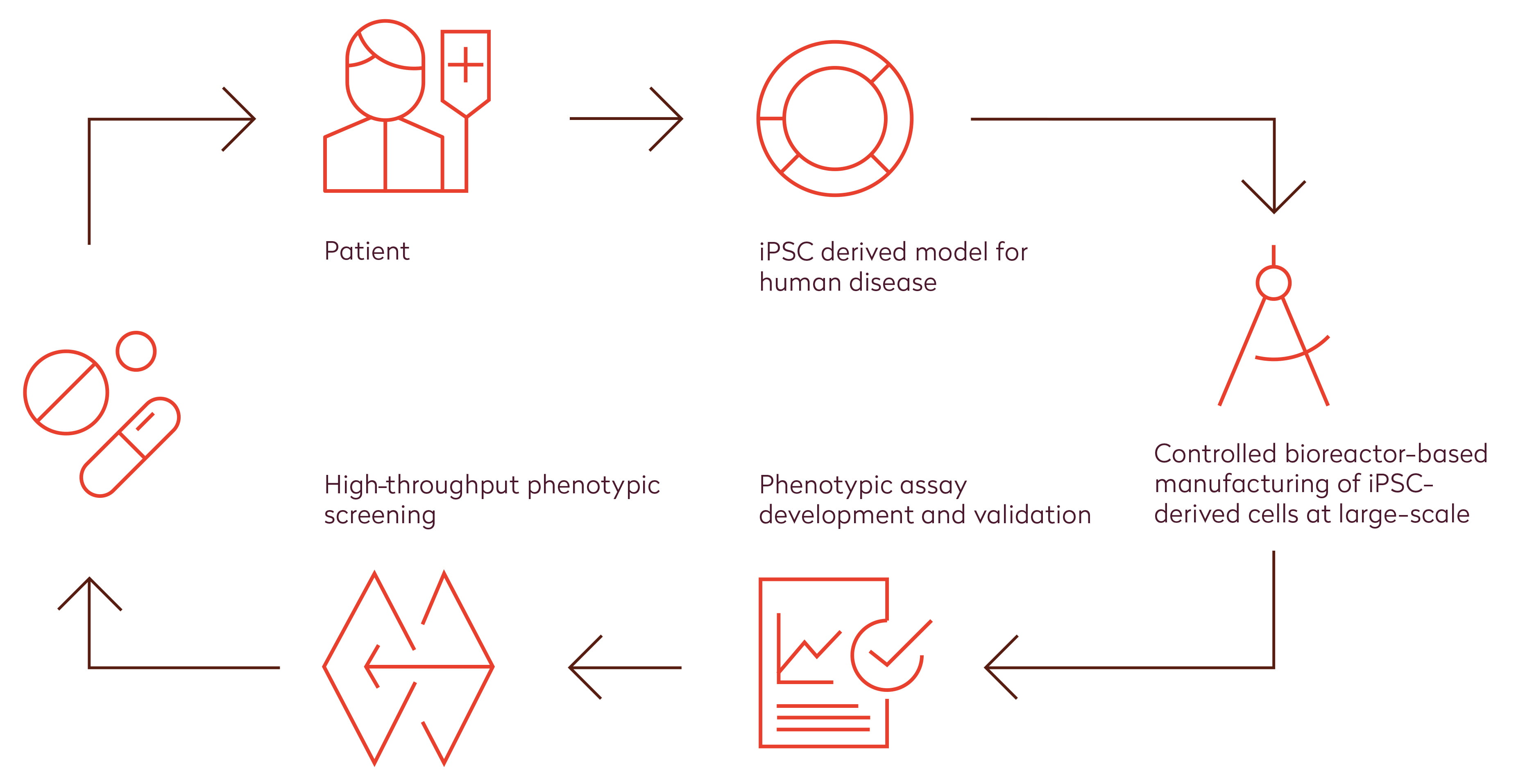

Figure 2: Human stem cell technology enables an end-to-end solution for efficient high-throughput phenotypic screening in drug discovery and development. State-of-the-art hiPSC technologies allow us to generate relevant human cell models for disease derived from virtually any patient. With the use of controlled bioreactors, high quality cells can be manufactured in large quantities. Combined with excellent capabilities on assay development, this enables high-throughput screening of small molecule libraries using a phenotypic approach. Altogether, this holds great promise for very reliable efficacy testing of novel drugs at a high throughput during early stages of drug discovery and development.
Controlled bioreactor-based large-scale manufacturing of hiPSC-based cardiomyocytes
Reproducible large-scale manufacturing of hiPSC-derived functional human cells of high quality, and implementation in automated systems for high-throughput screening is integral to implementing hiPSC-based disease models in drug discovery and development. To enable robust and scalable production of our cells for HTS applications, we have implemented a bioprocessing pipeline comprising state-of-the-art bioreactor systems, which allows us to optimise processes at 15mL scale to validate the most promising conditions at mid-scale of 100-250mL and to provide confidence to bring the process to the necessary 1-10L scale. Using a quality by design approach,8 we developed a robust and controlled process for large-scale manufacturing of high quality iPSC-derived cardiomyocytes. The resulting process is serum-free and provides a relatively mature cardiomyocyte model without genetic modifications, recapitulating a human cardiomyocyte’s contractile and electrophysiological profile with high predictivity in our cardiac toxicity and efficacy assays.9 The quality of cells was confirmed by the presence of cardiomyocyte markers (cTNT and MLC2v) in multiple batches and further corroborated by the electrophysiological properties (including sodium spike amplitude, field potential duration and beat rate) in multi-electrode arrays (MEA), demonstrating the relatively mature phenotype of the cells. The expected response to cardioactive reference compounds further confirmed the functionality of bioreactor-derived cardiomyocytes.
Development of robust cell-based phenotypic assays to assess cardiac functionality
Previously, we developed various functional applications dedicated for phenotypic screening.9 Some of these assays are high-throughout compatible, eg, high-content imaging or biochemical assays, which are being used in primary compound screenings. Other assays are low/medium throughput, such as contractility and electrophysiology assays, which are being used as secondary assays for hit confirmation. A successful drug efficacy screening study requires not only a physiologically-relevant cell model, but also a predictive and validated assay to test the effect of drug candidates. The assay performance is evaluated using putative quality-controlled criteria such as S/B ratio, Z-prime and %CV to ensure robustness of high-throughput compound screenings, with an aim to develop highly robust assays in a complex cell-based platform.
High-throughput efficacy screen in a diseased model of hypertrophy
To demonstrate the actual application of the bioreactor-derived cardiomyocytes in HTS we selected a model of hypertrophic cardiomyopathy (HCM). HCM is an autosomal dominant disease of the cardiac sarcomere and is estimated to be the most prevalent hereditary heart condition in the world.10 HCM causes abnormal thickening of the left ventricular myocardium, and results in elevated risk for various clinical complications such as progressive heart failure, arrhythmia, and sudden cardiac death. A peptide fragment NT-proBNP can be measured in patient serum and is established as a clinical biomarker for the diagnosis of hypertrophy. To build a HTS-relevant assay we took advantage of the fact that endothelin-1 (ET-1, a potent vasoconstrictor peptide produced by endothelial and smooth muscle cells) can induce hypertrophy in cardiomyocytes.11 We assessed the hypertrophic phenotype of the cells by measuring NT-proBNP secretion in the presence of ET-1 and various test compounds. The developed assay is fully automated using Fluent workstation starting from cell culturing in 384-well plates until last steps including automated data analysis. The assay is robust (S/B > 2) for both ET-1 induction and inhibition of the hypertrophy with our positive control, and also reproducible (%CV < 20 for vehicle and positive controls, and Z-factor=0.4). Screening statistics from duplicate plates once more confirmed the reproducibility of the assays (Pearson Correlation > 0.6). Taken together, this demonstrates the successful implementation of iPSC-derived cardiomyocytes in a high-throughput screening campaign.
Conclusion
Altogether, with the current results we demonstrate an end-to-end solution for more efficient and cost-effective drug discovery using a combination of cutting-edge hiPSC, bioprocessing, and high-throughput screening technologies. Human-induced stem-cell-based cellular models in combination with automated phenotypic screening now enables reliable efficacy testing of drugs at a high throughput, thereby revolutionising the drug discovery and development process. Phenotypic screening based on human models not only reduces the number of late-stage compound failures, it also allows continuation of projects that would otherwise fail. Most importantly, the state-of-the-art technologies discussed in the current review hold great promise to get more drugs to patients faster.
Acknowledgements
The authors have received funding from the European Union’s Horizon 2020 research and innovation programme under grant agreement No 726513. The authors are grateful to all the members of Ncardia’s disease modelling (Dr Oscar Bartulos) and bioprocess development (Dr Yang Jiang) teams for their expertise and gracious support.
Biographies








References
- Scannell JW, Blanckley A, Boldon H, Warrington B. Diagnosing the decline in pharmaceutical R&D efficiency. Nat Rev Drug Discov. 2012 Mar 1;11(3):191-200.
- Pammolli F, Magazzini L, Riccaboni M. The productivity crisis in pharmaceutical R&D. Nat Rev Drug Discov. 2011 Jun;10(6):428-38.
- DiMasi JA, Grabowski HG, Hansen RW. Innovation in the pharmaceutical industry: New estimates of R&D costs. J Health Econ. 2016 May;47:20-33.
- Vincent F, Loria P, Pregel M, Stanton R, Kitching L, Nocka K, Doyonnas R, Steppan C, Gilbert A, Schroeter T, Peakman MC. Developing predictive assays: the phenotypic screening “rule of 3”. Sci Transl Med. 2015 Jun 24;7(293):293ps15.
- Haasen D, Schopfer U, Antczak C, Guy C, Fuchs F, Selzer How Phenotypic Screening Influenced Drug Discovery: Lessons from Five Years of Practice. Assay Drug Dev Technol. 2017 Aug/Sep;15(6):239-246. doi: 10.1089/adt.2017.796. Epub 2017 Aug 11. Review.
- Swinney DC, Anthony J. How were new medicines discovered? Nat Rev Drug Discov. 2011 Jun 24;10(7):507-19.
- Mullard Stem-cell discovery platforms yield first clinical candidates. Nat Rev Drug Discov. 2015 Sep;14(9):589-91.
- Lipsitz YY, Timmins NE, Zandstra Quality cell therapy manufacturing by design. Nat Biotechnol. 2016 Apr;34(4):393-400.
- Mulder P, de Korte T, Dragicevic E, Kraushaar U, Printemps R, Vlaming MLH, Braam SR, Valentin JP. Predicting cardiac safety using human induced pluripotent stem cell-derived cardiomyocytes combined with multi-electrode array (MEA) technology: A conference report. J Pharmacol Toxicol Methods. 2018 May – Jun;91:36-42.
- Maron BJ, Gross BW, Stark SI. Images in cardiovascular medicine. Extreme left ventricular hypertrophy. Circulation. 1995 Nov 1;92(9):2748.
- Kuwahara K, Saito Y, Nakagawa O, Kishimoto I, Harada M, Ogawa E, Miyamoto Y, Hamanaka I, Kajiyama N, Takahashi N, Izumi T, Kawakami R, Tamura N, Ogawa Y, Nakao K. The effects of the selective ROCK inhibitor, Y27632, on ET-1-induced hypertrophic response in neonatal rat cardiac myocytes – possible involvement of Rho/ROCK pathway in cardiac muscle cell hypertrophy. FEBS Lett. 1999 Jun 11;452(3):314-8.
- Davis RP, van den Berg CW, Casini S, Braam SR, Mummery CL. Pluripotent stem cell models of cardiac disease and their implication for drug discovery and development. Trends Mol Med. 2011 Sep;17(9):475-84.
The rest of this content is restricted - login or subscribe free to access


Why subscribe? Join our growing community of thousands of industry professionals and gain access to:
- quarterly issues in print and/or digital format
- case studies, whitepapers, webinars and industry-leading content
- breaking news and features
- our extensive online archive of thousands of articles and years of past issues
- ...And it's all free!
Click here to Subscribe today Login here
Related organisations
Ncardia



