Gene differences that impact response to Cryptococcus identified
Posted: 17 July 2019 | Drug Target Review | No comments yet
Using human data, a new study has examined how Cryptococcus genes impact patients with Cryptococcus Neoformans.
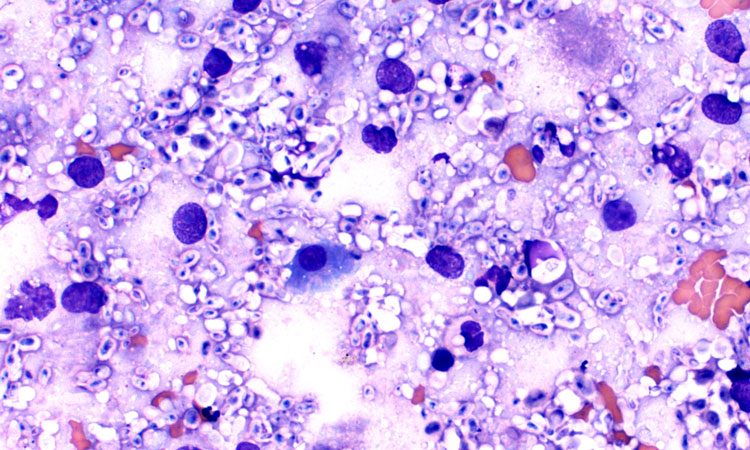

A new study by Kirsten Nielsen, PhD, Professor, Department of Microbiology and Immunology, University of Minnesota, Medical School and colleagues has examined how Cryptococcus genes impact patients with Cryptococcus Neoformans, using human data.
Nielsen’s last study found that the pathogen was driving the outcome of the Cryptococcus infection and this study goes on to examine the underlying genetic differences.
“We looked at differences in disease between patients – whether the patient lived or died, how the patient’s immune system responded to the infection, and whether the antifungal drug treatment worked well – and we asked: ‘How do genetic differences in the Cryptococcus strains impact the disease variables?'” explained Nielsen.
Drug Target Review has just announced the launch of its NEW and EXCLUSIVE report examining the evolution of AI and informatics in drug discovery and development.
In this 63 page in-depth report, experts and researchers explore the key benefits of AI and informatics processes, reveal where the challenges lie for the implementation of AI and how they see the use of these technologies streamlining workflows in the future.
Also featured are exclusive interviews with leading scientists from AstraZeneca, Auransa, PolarisQB and Chalmers University of Technology.
The study found that there are 40 genes that are crucial to the ability of Cryptococcus to change the outcome of human disease, which have never before been identified as important: “We can take this new information generated using the human data and show how the genes work in other models,” Nielsen continued. “When we deleted the genes, it changed the ability of Cryptococcus to cause disease in a model system, so we know that they are important in disease.”
Nielsen and her colleagues hope that identifying which versions of genes are important for patient survival will ultimately lead to better treatment of patients.
“We hope that this will have clinical benefits in the future. If we can figure out why certain strains are more deadly, and identify which patients have those strains, we can treat them differently. This will hopefully decrease reliance on toxic antifungals,” added Katrina Jackson, a graduate student from the University of Minnesota Medical School, who was involved in the project.
The study, ‘Identification Of Pathogen Genomic Differences That Impact Human Immune Response And Disease during Cryptococcus neoformans Infection’ was published in the journal MBio by American Society for Microbiology.
Related topics
Analysis, Disease research, Gene testing, Research & Development
Related conditions
Cryptococcus Neoformans
Related organisations
University of Minnesota
Related people
Kirsten Nielsen PhD



